Cladistics looks for the relationships between taxa in terms of an evolutionary cost. By minimizing this cost, it is expected to find an evolutionary scenario that closely matches the hierarchical diversification process through transmission with modification. In other words, inheritance of innovations from common ancestors is the simplest way to explain diversity.
To understand why this can only be achieved by using character (or parameter) matrix instead of distances that measures global similarities, consider the case of a journey between two cities.
Looking at a map, you can easily measure the distance between the two cities with a rule. This might be ok if you fly, but this is not very useful if you travel by car because then you have to take into account the landscape and the existing roads among other things.
First you have to look at a precise roadmap, and compute for each road and considering all possible bifurcations, the true number of kilometers you will have to travel. Note that there is no trick to avoid looking at all possible paths. Then you can decide to choose the shortest way according to the parsimony criterion.
But cost might not be measured by kilometers only. You may consider the time it takes. Highways are certainly faster, but you should consider the probability of traffic jams or that of slow trucks or animals on smaller roads. It is common wisdom that the quickest ways are not necessarily the shortest or the most direct ones.
You can also think about money with the fuel you will burn. Depending on your car, depending on the slopes for the different roads, the cost can be quite different in every case.
Lastly, you can consider the pleasure or the comfort of the journey. This is certainly less quantitative and objective, but still important.
We have here a typical multivariate problem, and defining an evolutionary cost is not always straightforward. As the above should illustrate, character-based methods like cladistics explore an unknown landscape with a metrics which is defined by the choice of the multivariate cost. Indeed, for living organisms or for galaxies, there is no roadmap…
Distance-based approaches assume a metrics and do not care very much on the cost (and even on the landscape). To understand further the difference, let us be more precise and consider the following parameter or character matrix:
| p1 | p2 | p3 | p4 | |
| O | 0 | 0 | 0 | 0 |
| A | 1 | 0 | 0 | 0 |
| B | 0 | 1 | 1 | 0 |
| C | 0 | 1 | 1 | 1 |
If you have learned to build a tree, you are able to find that the most parsimonious tree rooted with O is:
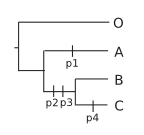 Now, from the parameters, you can compute a distance, the most common being the euclidian distance. The corresponding distance matrix (showing here the square of the euclidian distance) is:
Now, from the parameters, you can compute a distance, the most common being the euclidian distance. The corresponding distance matrix (showing here the square of the euclidian distance) is:
| O | A | B | C | |
| O | 0 | 1 | 2 | 3 |
| A | 1 | 0 | 3 | 4 |
| B | 2 | 3 | 0 | 1 |
| C | 3 | 4 | 1 | 0 |
One could also compute the “edit” or Levenshtein distance, which measures the number of substitution (here 0-1) occuring in the full set of parameters between two objects. The matrix distance in the present case is identical to the one above. Note that even though it might look like cladistics because it compares the changes in parameter values, it is a distance and thus measures these changes globally.
From any character matrix you can compute a distance matrix, but the reverse is most generally untrue. Hence, somehow, when we use distances, we loose some information.
From a distance matrix, we can build a hierarchical tree representing the relative distances between the objects. Using hclust in R, this gives:
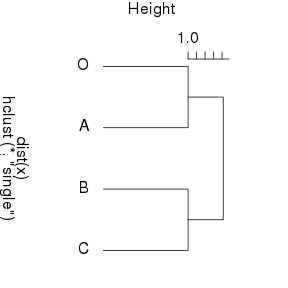 From this tree, one concludes that there are two groups: (O,A) and (B,C). The distance within the two members of each group is 1 while the minimum distance between the groups is 2. So the two methods agree that B and C are very close to each other and could form a group, but cladistics do not see any reason to put O and A in a same group. Indeed the cladogram is easier to interpret because it must be read in terms of the evolutionary cost.
From this tree, one concludes that there are two groups: (O,A) and (B,C). The distance within the two members of each group is 1 while the minimum distance between the groups is 2. So the two methods agree that B and C are very close to each other and could form a group, but cladistics do not see any reason to put O and A in a same group. Indeed the cladogram is easier to interpret because it must be read in terms of the evolutionary cost.
In general, it appears that character-based and distance-based analysis in phylogeny give very close results. I think this is because if only synapomorphies are used, which ideally should be the case for cladistics, then the landscape is not too much tortuous so that the metrics assumed by distance-based approaches is more or less adequate.

#1 by dranorter on July 3, 2013 - 07:10
You say there is no trick to avoid looking at all possible paths. However, I think you are thinking of the Traveling Salesman Problem, where we must visit every node in an efficient manner. When traveling from point A to point B there certainly are useful tricks and shortcuts. Dijkstra’s Algorithm is a common approach, and is much faster than looking at all possibilities.
Of course, the ‘multivariate’ case gets more complicated. I’m only disagreeing with that one sentence.